TESTING AND ANALYSIS
By Simerdeep Kaur, Ph.D., Post-Doctoral Research Fellow, Oliver Lab, Department of Food Science, Purdue University
When Dead Bacteria Trigger False Alarms: Rethinking a Common "Live-Dead" Test
False positives may result from PMA use in PCR testing, which can lead to food recalls and waste
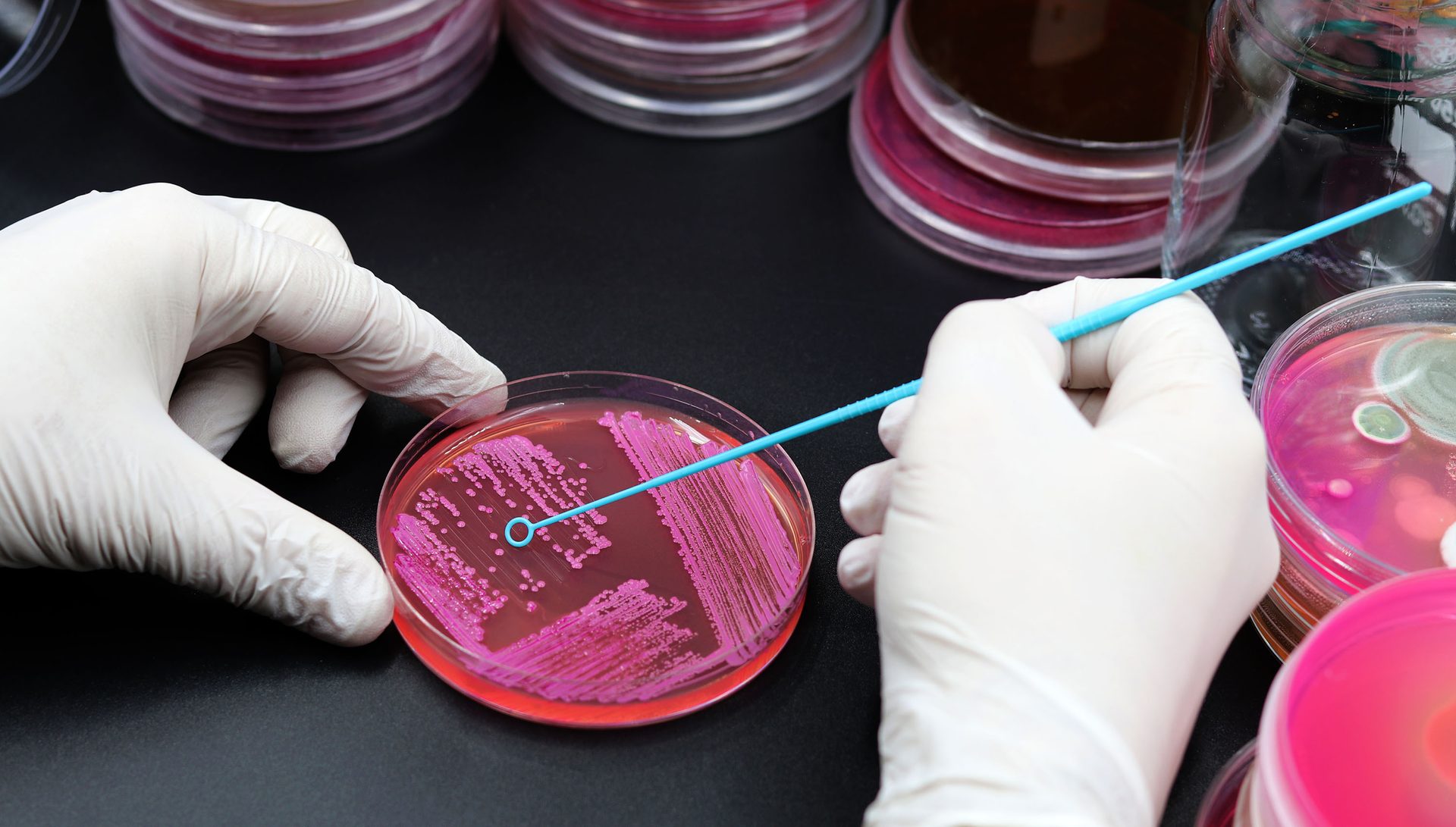
Image credit: TopMicrobialStock/iStock/Getty Images Plus via Getty Images
SCROLL DOWN
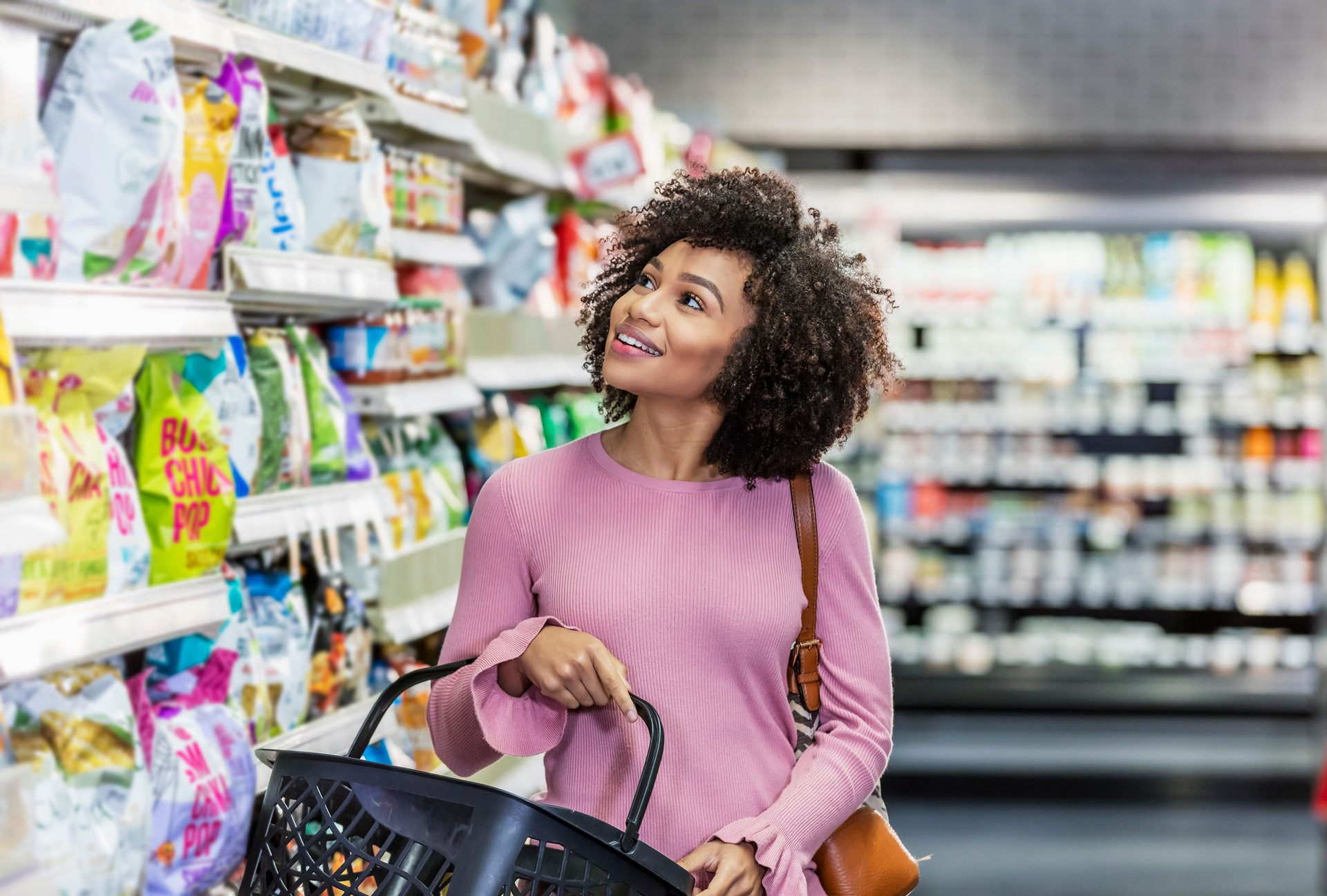
Every year, food products are pulled from shelves due to safety recalls. These recalls happen for many reasons, from unlabeled allergens to detected pathogen contamination in food processing facilities.1 Recalls are an expensive and wasteful affair: a single recall can cost a company over $10 million in direct costs alone.2 Beyond the dollars, there is the staggering waste of food. In just the first half of 2024, the U.S. Food and Drug Administration (FDA) ordered the destruction of nearly 85 million units of food over safety concerns.3 Often, all that food ends up in landfills, even if it might have been safe to eat.
Foodborne pathogens can be detected either after an outbreak has occurred or, ideally, before contaminated products reach consumers. In this preventive approach, companies err on the side of caution. If a lab test suggests that a pathogen might be present in a product, the only safe move is to issue a recall and throw out the product. But what if some of those positive test results are false alarms?
DNA Tests: Powerful Tools with a Limitation
Today's food safety labs rely on nucleic acid amplification tests (NAATs) to detect the DNA of foodborne pathogens. Techniques like polymerase chain reaction (PCR) and loop-mediated isothermal amplification (LAMP) can detect pathogens with remarkable speed and sensitivity. These DNA-based assays have revolutionized food safety testing by allowing the detection of pathogens before they cause illness, or tracing the source of contamination after an outbreak. This powerful tool has a notable limitation, however: DNA can long outlive the bacteria it came from.
When bacteria die, their DNA can remain intact for days or even weeks. So, if a facility had a contamination that was properly remediated and the bacteria were killed, traces of their DNA might still linger. A PCR test swab of the area could be positive, detecting dead bacteria. This means that positive PCR results can sometimes be false alarms.
For food companies and safety regulators, this limitation poses a dilemma. In many workflows, NAAT positives are treated as presumptive and are followed by confirmatory testing (often culture isolation) before final disposition. However, if a positive signal is driven by dead-cell DNA (especially in environmental follow-up testing), then the product hold, investigation, and sometimes the destruction of food that follows might be unnecessary. This challenge has motivated scientists to find ways to make DNA-based tests more discerning, so they respond only to live bacteria.
A Dye to Tell "Dead" from "Alive"
About 15–20 years ago, scientists developed a clever workaround to PCR's live/dead problem by using a dye called propidium monoazide (PMA). PMA is a molecule designed to distinguish between live and dead cells in PCR-based assays. When added to a sample, PMA can enter bacteria that have compromised cell membranes and bind to their DNA. After an activation step with blue light, the PMA cross-links to the DNA, locking it away so it cannot be amplified by PCR. Meanwhile, bacteria with intact cell membranes are meant to be spared. In theory, the result is a "viability PCR": only living bacteria yield a positive signal, because DNA from dead cells has been cross-linked by the dye, eliminating false positives.
Since its introduction in the early 2000s, PMA-based viability PCR has been widely adopted in research and testing. In food safety, it has been used to refine PCR tests to minimize false positives due to dead cells. It has also been applied in microbiome research (like 16S rRNA gene sequencing) to avoid over-counting bacteria that are no longer alive.
Over the years, scattered reports began to suggest that the PMA method was not foolproof. Some studies observed that PMA treatment can interfere with detecting live cells. At the same time, other reports noted that PMA sometimes fails to fully eliminate the signal from dead cells. These concerns were anecdotal or limited to specific conditions, and many labs continued to use PMA, assuming that with the right protocol it would reliably reflect microbial viability.
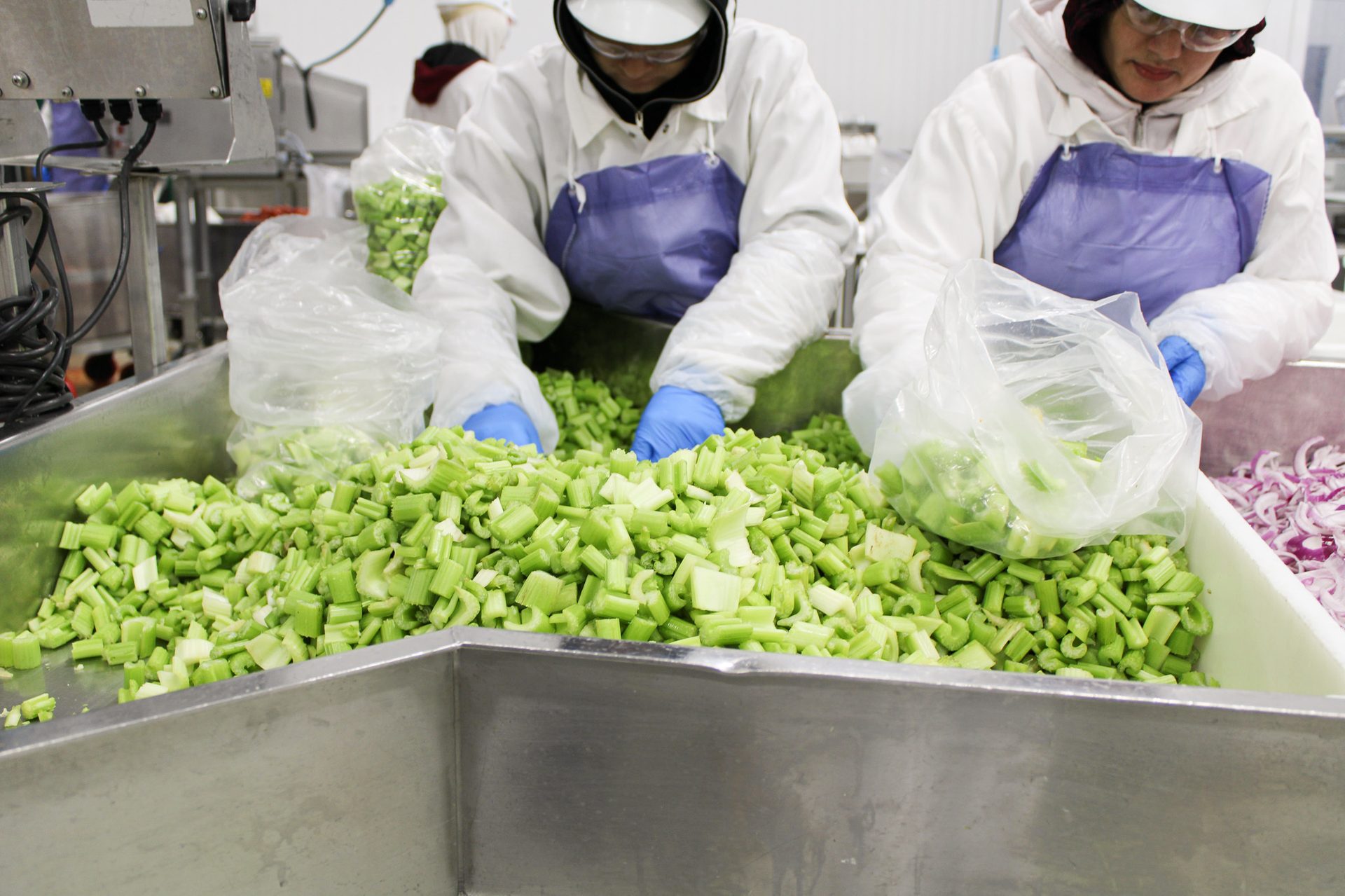
FIGURE 1. Cut Fresh has a number of best practices, auditing, and certification efforts in place to ensure the highest standards of food safety, food quality, and sanitation (Image courtesy of Cut Fresh LLC / Anna Patsakham)

“[PMA] is suitable only in carefully controlled cases—essentially, if a rough estimate is known of how many dead cells are present—which defeats the purpose of using it in unknown samples.”
Unreliable Results: What New Research Found
Unfortunately, real-world biology is messier than the theory. As I found in our recent research, PMA does not always behave so cleanly. An overall conclusion of the study was that PMA-based viability assays can be unreliable and inconsistent, especially when the number of dead cells versus live cells in a sample is unknown.
PMA is supposed to prevent DNA from dead bacteria from showing up in test results. However, our researchers found that this only works under very specific conditions. If there are too many dead cells, then there is not enough PMA to block all the leftover DNA, and dead bacteria still show up as a positive result. On the flip side, if there are mostly live cells and too much PMA is used, it can interfere with those living bacteria, making it harder to detect a real contamination. That means labs must hit a very narrow "just right" window. This is tricky, because in real food samples, the number of live or dead bacteria is rarely known.
It gets more complicated. The research team tested how PMA works on different types of bacteria that are often responsible for foodborne illness: E. coli, Salmonella, and Listeria. The dye did not affect each pathogen in the same way. In some bacteria, even the live cells were sensitive to the dye and gave off a weaker signal in tests. Others were not affected as much. What this means is that a PMA treatment that works well for Salmonella might undercount live E. coli, or vice versa.
The test's efficiency also matters. The researchers compared two common methods: a highly sensitive test (qPCR) and a less sensitive one (LAMP). PMA seemed to work better in the less sensitive test, but only because the test could not detect leftover DNA that was still slipping through. With the more sensitive method, the flaws in PMA became more obvious.
The research report plainly states that PMA is unreliable for viability assays when the concentration and composition of the bacterial mixtures are unknown. It is suitable only in carefully controlled cases—essentially, if a rough estimate is known of how many dead cells are present—which defeats the purpose of using it in unknown samples.
Implications for Food Safety and Future Approaches
The findings carry important implications for food safety testing. For food producers and testing labs, it suggests that relying on PMA-PCR alone to confirm a pathogen's viability could be problematic. An assay that is meant to prevent false positives might instead be introducing new uncertainties.
In the bigger picture, the study underscores a need for better methods to differentiate live and dead pathogens in rapid testing. PMA is not the only approach out there. Researchers are exploring other options—for example, targeting RNA (which degrades much faster after cell death than DNA) as an alternative strategy.4 Each alternative approach has its pros and cons, and none is a perfect solution. There is also interest in developing entirely new kinds of sensors—for example, detecting metabolic activity of bacteria as proof of life or using advanced techniques like microfluidics.5 Some testing workflows have even added extra steps to try to mitigate false positives—e.g., by physically removing or "cleaning" residual DNA from dead cells before running the PCR test.6 The challenge is significant: the method must be both reliable and practical. It should integrate into routine testing without too much extra cost or complexity, because food companies need fast, affordable tests to keep up with high production volumes and rigid safety standards.
The consequences of false positives are not just economic; they also affect how we perceive food safety. Frequent recalls based on precaution can erode public trust in the food supply and in testing methods themselves. On the flip side, a truly reliable viability assay would empower producers to make more nuanced decisions and increase their frequency of testing. Given that an estimated 2.4 percent of U.S. food waste is related to safety concerns like recalls,3 even modest improvements in test accuracy could translate into saving millions of pounds of food from the garbage heap each year.
References
- Morawicki, R. "Food Recalls: An Unnecessary and Preventable Factor in Food Waste." Journal of Agriculture, Food Systems, and Community Development 14, no. 1 (2025). https://www.foodsystemsjournal.org/index.php/fsj/article/view/1318.
- Skip Shapiro Enterprises. "The Real Cost of Food Recalls: How to Minimize It?" Food Solutions Blog. May 26, 2023. https://shapiroe.com/blog/the-cost-of-food-recalls/.
- Nemo, L. "USDA and FDA food recalls are on the rise. What happens to the waste?" WasteDive. October 14, 2025. https://www.wastedive.com/news/food-recall-organic-waste-usda-fda-misfits-markets/761320/.
- Liu, Y., C. Wei, H. Wan, et al. "Preliminary Study on Rapid and Simultaneous Detection of Viable Escherichia coli O157:H7, Staphylococcus Aureus, and Salmonella by PMA-mPCR in Food." Molecules 28, no. 15 (August 2023). https://www.mdpi.com/1420-3049/28/15/5835.
- Santander, R.D., C.L. Meredith, and S.G. Aćimović. "Development of a Viability Digital PCR Protocol for the Selective Detection and Quantification of Live Erwinia amylovora Cells in Cankers." Scientific Reports 9 (August 2019). https://www.nature.com/articles/s41598-019-47976-x.
- Liu, J., I. Jasim, A. Abdullah, et al. "An Integrated Impedance Biosensor Platform for Detection of Pathogens in Poultry Products." Scientific Reports 8 (October 2018). https://www.nature.com/articles/s41598-018-33972-0.
Additional Reading
- Macher, J., Ed. Bioaerosols: Assessment and Control. American Conference of Governmental Industrial Hygienists, 1999.
- Lorenzo, J.M., P.E. Munekata, R. Dominguez, M. Pateiro, J.A. Saraiva, and D. Franco. "Chapter 3: Main Groups of Microorganisms of Relevance for Food Safety and Stability: General Aspects and Overall Description." Innovative Technologies for Food Preservation. Academic Press, 2018. https://www.sciencedirect.com/science/chapter/edited-volume/pii/B9780128110317000030.
- Fernstrom, A. and M. Goldblatt. "Aerobiology and Its Role in the Transmission of Infectious Diseases." Journal of Pathogens (January 2013): 493960. https://pmc.ncbi.nlm.nih.gov/articles/PMC3556854/.
Simerdeep Kaur, Ph.D. is a microbiologist advancing microbial food safety across agricultural and food systems through molecular detection, environmental data integration, and systems-level analysis. She received her M.Sc. degree in Microbiology from Panjab University in India and her Ph.D. in Agricultural and Biological Engineering from Purdue University. She is currently a postdoctoral researcher in Purdue University's Department of Food Science, investigating pathogen persistence, sanitation systems, and biofilm-associated survival in food environments.

